HistoCRF
Conditional Random Fields for Interactive Refinement of Histopathological Predictions [Submitted to ICASSP 2026]
Assisting pathologists in the analysis of histopathological images has high clinical value, as it supports cancer detection and staging. In this context, histology foundation models have recently emerged. Among them, Vision-Language Models (VLMs) provide strong yet imperfect zero-shot predictions. We propose to refine these predictions by adapting Conditional Random Fields (CRFs) to histopathological applications, requiring no additional model training.
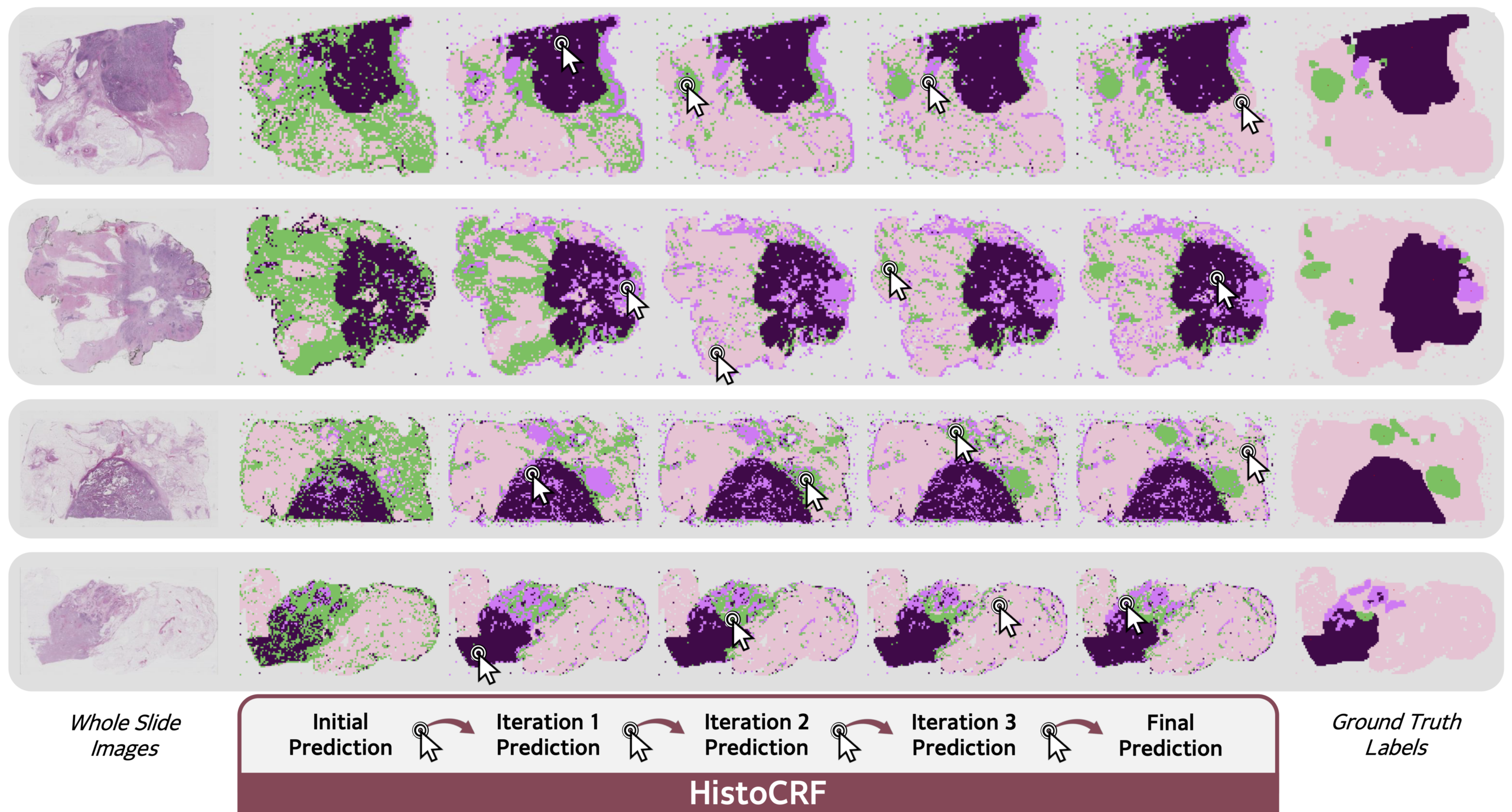
We present HistoCRF, a CRF-based framework, with a novel definition of the pairwise potential that promotes label diversity and leverages expert annotations. We consider three experiments: without annotations, with expert annotations, and with iterative human-in-the-loop annotations that progressively correct misclassified patches.
Experiments on five patch-level classification datasets covering different organs and diseases demonstrate average accuracy gains of 14.3\% without annotations and 27.1\% with only 100 annotations, compared to zero-shot predictions.
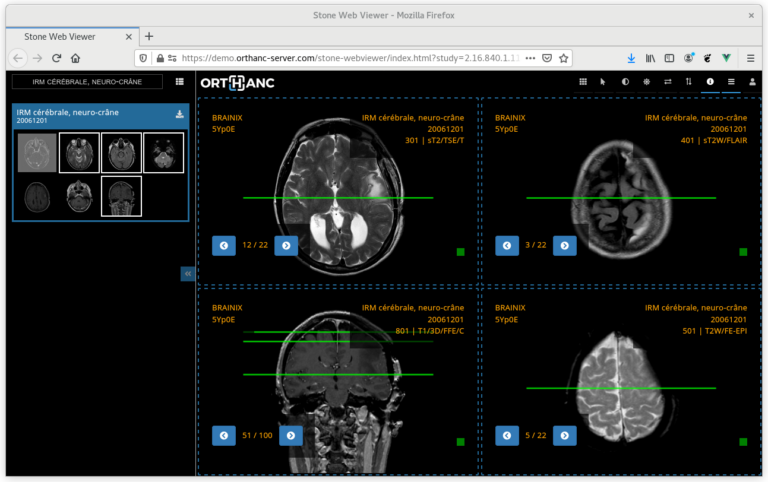
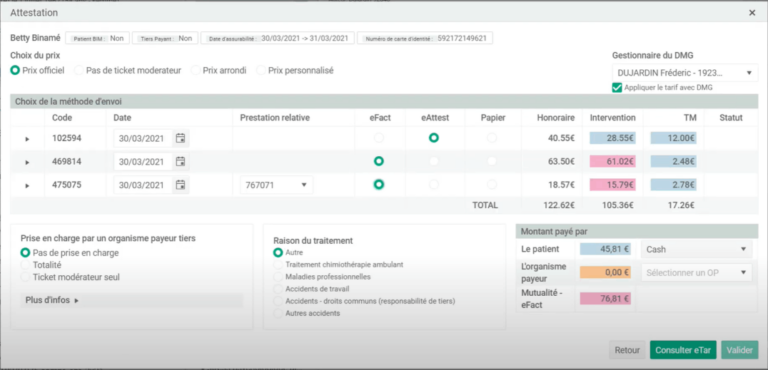
Moreover, integrating a human in the loop reaches a further gain of 32.7\% with the same number of annotations. An example of a single iterative message passing step is shown on the right.
Authors: T. Godelaine, M. Zanella, K. El Khoury, S. Mahmoudi, B. Macq, C. De Vleeschouwer
Contents
Installation
📌 NB: The Python version used is 3.10.4.
- Create a virtual environment
python3 -m venv histocrf_venv
source histocrf_venv/bin/activateClone the GitHub repository
pip3 install torch==2.5.1 torchaudio==2.5.1 torchvision==0.20.1
git clone https://github.com/tgodelaine/HistoCRF.gitInstall the required packages
cd HistoCRF
pip install -r requirements.txt- Models downloads:
| Model | 🔗 Download Link | Folder name |
|---|---|---|
| CONCH | 📥 Link | ./root/HistoCRF/CONCH |
| UNI2h | 📥 Link | No download is needed but you do need to request access. |
Important: For the model CONCH, do not forget to download the checkpoints.
- Datasets downloads:
| Dataset | 🔗 Download Link | Folder name |
|---|---|---|
| BACH | 📥 Link | ./data/thunder/datasets/BACH |
| BRACS | 📥 Link | ./data/BRACS |
| ESCA | 📥 Link | ./data/thunder/datasets/ESCA |
| NCT | 📥 Link | ./data/NCT-CRC-HE-100K |
| SICAP | 📥 Link | ./data/SICAP_MIL |
Important: Each dataset must be placed in the directory ./data/.
Usage
- Once the datasets are downloaded, you must extract the features of each patch in the dataset. By using the following command line, a .npz file will be created at
./root/data_processed.
cd HistoCRF
python3 extraction.py --root_dir ./root/ --data_dir ./data/ --model {model} --dataset {dataset}- Once the features are extracted, you can run the experiments using the following command lines. The results will be saved into a JSON file for further analysis at
./root/results.
| Experiment | Command line |
|---|---|
| LP | python3 inference_iteration_split_split.py --root_dir ./root/ --model {model} --dataset {dataset} --uni_pot softmax --weight 0.1 0.01 --pair_pot minus_model_features model_features_ann --compat indicator potts --sparse_method cossim --n_affinity 0 16 --n_annotations {n_annotations} --annotation_method error -ann_iterations {ann_iterations} --N_UP -100 --N_PROP 1 --seed {seed} |
| HistoCRF | python3 inference_iteration_split_split.py --root_dir ./root/ --model {model} --dataset {dataset} --uni_pot softmax --weight 0.1 0.01 --pair_pot minus_model_features model_features_ann --compat indicator potts --sparse_method cossim --n_affinity 0 16 --n_annotations {n_annotations} --annotation_method error -ann_iterations 1 --N_UP -1 --N_PROP 50 --seed {seed} |
| HistoCRF-HITL | python3 inference_iteration_split_split.py --root_dir ./root/ --model {model} --dataset {dataset} --uni_pot softmax --weight 0.1 0.01 --pair_pot minus_model_features model_features_ann --compat indicator potts --sparse_method cossim --n_affinity 0 16 --n_annotations {n_annotations} --annotation_method error -ann_iterations {ann_iterations} --N_UP -1 --N_PROP 50 --seed {seed} |
Contact
If you have any questions, you can contact us by email: tiffanie.godelaine@uclouvain.be